PAIRWISE ALIGNMENT TECHNIQUES
A manner of construction of dot plot matrix is shown below. Here for identical residue we mark it as a dot.
A pair of distantly related sequences with fewer similarities has a much noisier plot as shown above. Dot plot helps in comparison of sequences on the basis of evolutionary relation, structural similarity, and physiochemical properties etc.
Pairwise sequence alignments can only be used between two sequences at a time but they are very efficient to find out the similarities. Pairwise comparison is a fundamental process in sequence analysis, which seeks out relationships based on sequence properties. Database searching is used to find out the sequence similarity searches. Pairwise sequence alignment methods are used to find the best matching piecewise (local) or global alignments of a two query sequences. The three primary methods of producing pairwise alignments are dot-matrix methods, dynamic programming and word methods. But of these methods, dot-matrix method is the popular one for pairwise alignments, and for multiple alignments, the other two methods are commonly used.
DOT PLOT SEQUENCE ALIGNMENT:
The most basic method of comparing two sequence is a visual approach known a dot-plot. Dot lot is a biological sequence comparison plot. The dot-matrix approach is qualitative and simple.
A dot plot is a graphical method that allows the comparison of two biological sequences and identifies the regions of close similarity between them. The simplest way visualize the similarity between two protein sequences is to use a similarity matrix own as a dot-plot. From dot-plot it is easy to visually identify certain sequence features such as insertions, deletions repeats, inverted repeats etc.
There are two dimensional matrices, which have the sequences of the proteins being compared along the vertical and horizontal axes. To construct a dot-plot the two sequences are written along the top row and leftmost column of a two-dimensional matrix and a dot is placed at any point where the character in the appropriate columns matches. In some implementations the size or intensity of the dot is varied depending on the degree of similarity of the two characters, So the matrix sequence segments appear as runs of diagonal lines across the matrix. The dot plots can also be used to assess repetiveness in a single sequence.
A manner of construction of dot plot matrix is shown below. Here for identical residue we mark it as a dot.
 |
| Dot Plot |
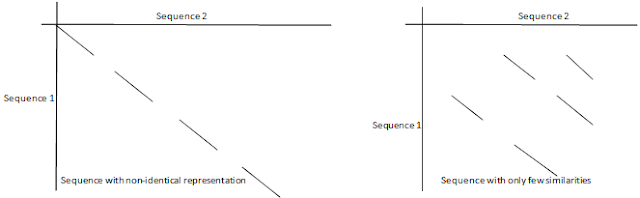
Within a dot plot two identical sequences are characterized by a single unbroken diagonal line across the plot as shown above. But two similar, but non-identical sequences will be characterized by a broken diagonal and here the interrupted region indicates the location of sequence mismatches.
A pair of distantly related sequences with fewer similarities has a much noisier plot as shown above. Dot plot helps in comparison of sequences on the basis of evolutionary relation, structural similarity, and physiochemical properties etc.